We
are interested in two broad applications of this technology in
C. elegans. First is the generation of recombinant 'zinc
finger nucleases' (ZFNs) that can be used to generate enzymes
that cause
breaks
in DNA at a specific site flanked by two component 9-mer recognition
sites:
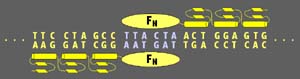
zinc finger
nuclease
In Drosophila,
the Carroll
lab (Utah) has shown that such breaks can lead to
mis-repair or repair from a transgene, allowing genes to be targeted
for
specific
changes. In C. elegans, similar breaks caused by transposable
elements can direct repair from transgenes. Our goal is to develop
a similar zinc-finger nuclease technology for C. elegans.
Our second
application is the generation of novel transcription factors.
Using 6-mer fingers attached to effector domains, such proteins
can be used to up- or down-regulate genes of interest. In C.
elegans, this could be used to study gene function in many
ways.
As a first
step towards developing this technology in C. elegans,
we have constructed a zinc finger library that allows generation
of multi-fingered coding regions using simple cloning. Our current
work is aimed at demonstrating function of ZFNs and expression
of chimeric transcription factors.
 Designer
Zinc Finger Proteins (Segal and Barbas, 2001) Designer
Zinc Finger Proteins (Segal and Barbas, 2001)
ZFNs
in Drosophila:  Bibikova et
al.,
2002 Bibikova et
al.,
2002  Bibikova et
al., 2003 Bibikova et
al., 2003
 Transgene
Repair in C. elegans
(Plasterk and Groenen, 1992) Transgene
Repair in C. elegans
(Plasterk and Groenen, 1992)
|